
We use molecular systematic tools to understand how lineages of plants are related to each other. Understanding genetic relationships among plant taxa (lineages, species, genera, families…) provides the basis for inferring all kinds of evolutionary processes at work in shaping the geographic distributions of plants (i.e., Where do they live now, in the past and in the future?) and their particular adaptations (i.e., How do they survive? How do their flowers function? What factors influence reproductive success?).
All of the methods rely on the polymerase chain reaction (PCR) to generate enough copies of DNA for analysis. Without PCR, we would have too few copies of the molecule to “read” genetic markers (DNA sequences, DNA fragment lengths, etc.). We use a few different types of genetic markers in our lab. These include…
RAD-Seq (Restriction Site Associated DNA) is a next-generation sequencing (NGS; high-throughput) sequencing method that has been extremely useful for elucidating gene flow patterns at the population level, either within a single species or among multiple species. We use RAD-Seq to understand gene flow in species complexes, groups of closely related species that (for a variety or reasons) maintain relative reproductive isolation. RAD Seq data are generated by digesting genomic DNA with restriction enzymes, selectively amplifying a subset of the fragments generated, and obtaining DNA data via next-generation sequencing of these fragments.
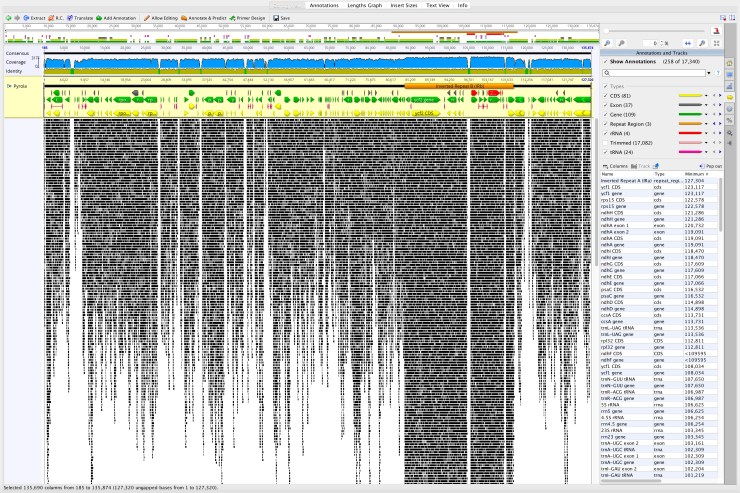
Genome skim is another NGS method for obtaining DNA sequences in a way that maximizes genomic breadth over depth in sampling coverage. It represents a relatively low-cost method for surveying the genomes, which is invaluable for studies of non-model organisms. Because the method can be optimized to achieve minimally sufficient sequencing depth while maximizing breadth, it is a reasonable method to use for our studies of species complexes, which require high sample sizes of individual plants.
AFLP (Amplified Fragment Length Polymorphism) is an older, restriction enzyme-based method for estimating gene flow and shares many similarities with RAD Sequencing with regard to the use of restriction digestion and selective amplification. However, the data output for AFLP are fragment lengths only. Yes, this means that homology assessment is based on fragment length. While the selective amplification steps help reassure us that fragments of the same length are homologous, the absence of sequence information is troubling to many. The use of AFLP for Dr. Jolles’ MS research provided the first DNA-based evidence that members of the P. picta species complex were indeed discrete taxa. Subsequent sequencing of a nuclear ribosomal DNA locus corroborated the AFLP data 🙂
Sanger sequencing was the primary method of generating DNA sequence data until several years ago, when NGS methods became more accessible to scientists, especially those with relatively low research budgets (like botanists, graduate students, etc). Sanger sequencing allows researchers to PCR amplify single genetic loci, ranging from ~100 to 5000 base pairs in length, for individual plant samples. Polymorphisms within these alignments (single sequence per plant sample included) are used to infer sister relationships and estimate phylogeny. Sanger sequencing is perfect for targeted sequencing of informative loci and is an affordable tool for conducting pilot studies.
- Nuclear loci are typically quickly evolving in plants and can be useful for evaluating recent speciation or lineage diversification. Many nuclear loci exist in multiple copy, making homology assessment and assumptions problematic. So-called “low copy” or single-copy nuclear loci can be identified using NGS, allowing for effective targeted sequencing of informative loci.
- Plastid loci are typically (but not always!) more slowly evolving in plants and can be useful for evaluating diversification at deeper nodes in a phylogeny (for example, the divergences among genera, families, and orders of plants). The plastid genome is typically inherited from a single parent, presenting some cool opportunities to follow maternal lineages. However, uniparental inheritance should be verified, not presumed!
